How to Add Factors to Continuous Variable in R Dplyr by Mean
Compute and Add new Variables to a Data Frame in R
This tutorial describes how to compute and add new variables to a data frame in R. You will learn the following R functions from the dplyr R package:
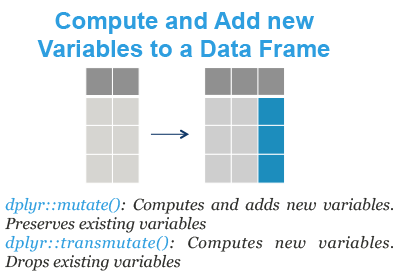
- mutate(): compute and add new variables into a data table. It preserves existing variables.
- transmute(): compute new columns but drop existing variables.
We'll also present three variants of mutate() and transmute() to modify multiple columns at once:
- mutate_all() / transmute_all(): apply a function to every columns in the data frame.
- mutate_at() / transmute_at(): apply a function to specific columns selected with a character vector
- mutate_if() / transmute_if(): apply a function to columns selected with a predicate function that returns TRUE.
Contents:
- Required packages
- Demo dataset
- mutate: Add new variables by preserving existing ones
- transmute: Make new variables by dropping existing ones
- Modify multiple columns at once
- Transform all column values
- Transform specific columns
- Summary
Required packages
Load the tidyverse packages, which include dplyr:
library(tidyverse) Demo dataset
We'll use the R built-in iris data set, which we start by converting into a tibble data frame (tbl_df) for easier data analysis.
my_data <- as_tibble(iris) my_data ## # A tibble: 150 x 5 ## Sepal.Length Sepal.Width Petal.Length Petal.Width Species ## <dbl> <dbl> <dbl> <dbl> <fct> ## 1 5.1 3.5 1.4 0.2 setosa ## 2 4.9 3 1.4 0.2 setosa ## 3 4.7 3.2 1.3 0.2 setosa ## 4 4.6 3.1 1.5 0.2 setosa ## 5 5 3.6 1.4 0.2 setosa ## 6 5.4 3.9 1.7 0.4 setosa ## # ... with 144 more rows mutate: Add new variables by preserving existing ones
Add new columns (sepal_by_petal_*) by preserving existing ones:
my_data %>% mutate(sepal_by_petal_l = Sepal.Length/Petal.Length) ## # A tibble: 150 x 6 ## Sepal.Length Sepal.Width Petal.Length Petal.Width Species ## <dbl> <dbl> <dbl> <dbl> <fct> ## 1 5.1 3.5 1.4 0.2 setosa ## 2 4.9 3 1.4 0.2 setosa ## 3 4.7 3.2 1.3 0.2 setosa ## 4 4.6 3.1 1.5 0.2 setosa ## 5 5 3.6 1.4 0.2 setosa ## 6 5.4 3.9 1.7 0.4 setosa ## # ... with 144 more rows, and 1 more variable: sepal_by_petal_l <dbl> transmute: Make new variables by dropping existing ones
Add new columns (sepal_by_petal_*) by dropping existing ones:
my_data %>% transmute( sepal_by_petal_l = Sepal.Length/Petal.Length, sepal_by_petal_w = Sepal.Width/Petal.Width ) ## # A tibble: 150 x 2 ## sepal_by_petal_l sepal_by_petal_w ## <dbl> <dbl> ## 1 3.64 17.5 ## 2 3.5 15 ## 3 3.62 16 ## 4 3.07 15.5 ## 5 3.57 18 ## 6 3.18 9.75 ## # ... with 144 more rows Modify multiple columns at once
We start by creating a demo data set, my_data2, which contains only numeric columns. To do so, we'll remove the column Species as follow:
my_data2 <- my_data %>% select(-Species) The functions mutate_all() / transmute_all(), mutate_at() / transmute_at() and mutate_if() / transmute_if() can be used to modify multiple columns at once.
The simplified formats are as follow:
# Mutate variants mutate_all(.tbl, .funs, ...) mutate_if(.tbl, .predicate, .funs, ...) mutate_at(.tbl, .vars, .funs, ...) # Transmute variants transmute_all(.tbl, .funs, ...) transmute_if(.tbl, .predicate, .funs, ...) transmute_at(.tbl, .vars, .funs, ...) - .tbl: a tbl data frame
- .funs: List of function calls generated by
funs(), or a character vector of function names, or simply a function. - …: Additional arguments for the function calls in .funs.
- .predicate: A predicate function to be applied to the columns or a logical vector. The variables for which .predicate is or returns TRUE are selected.
In the following sections, we'll present only the variants of mutate(). The transmute() variants can be used similarly.
Transform all column values
- Divide all columns value by 2.54:
my_data2 %>% mutate_all(funs(./2.54)) ## # A tibble: 150 x 4 ## Sepal.Length Sepal.Width Petal.Length Petal.Width ## <dbl> <dbl> <dbl> <dbl> ## 1 2.01 1.38 0.551 0.0787 ## 2 1.93 1.18 0.551 0.0787 ## 3 1.85 1.26 0.512 0.0787 ## 4 1.81 1.22 0.591 0.0787 ## 5 1.97 1.42 0.551 0.0787 ## 6 2.13 1.54 0.669 0.157 ## # ... with 144 more rows Note that, the dot "." represents any variables
- Function names will be appended to column names if
.funshas names or multiple inputs:
my_data2 %>% mutate_all(funs(cm = ./2.54)) ## # A tibble: 150 x 8 ## Sepal.Length Sepal.Width Petal.Length Petal.Width Sepal.Length_cm ## <dbl> <dbl> <dbl> <dbl> <dbl> ## 1 5.1 3.5 1.4 0.2 2.01 ## 2 4.9 3 1.4 0.2 1.93 ## 3 4.7 3.2 1.3 0.2 1.85 ## 4 4.6 3.1 1.5 0.2 1.81 ## 5 5 3.6 1.4 0.2 1.97 ## 6 5.4 3.9 1.7 0.4 2.13 ## # ... with 144 more rows, and 3 more variables: Sepal.Width_cm <dbl>, ## # Petal.Length_cm <dbl>, Petal.Width_cm <dbl> Note that, the output variable name now includes the function name.
Transform specific columns
- mutate_at(): transform specific columns selected by names:
my_data2 %>% mutate_at( c("Sepal.Length", "Petal.Width"), funs(cm = ./2.54) ) - mutate_if(): transform specific columns selected by a predicate function.
mutate_if() is particularly useful for transforming variables from one type to another.
my_data %>% mutate_if(is.factor, as.character) Round all numeric variables:
my_data %>% mutate_if(is.numeric, round, digits = 0) Summary
This article describe how to add new variable columns into a data frame using the dplyr functions: mutate(), transmute() and variants.
-
mutate(iris, sepal = 2*Sepal.Length): Computes and appends new variable(s). -
transmute(iris, sepal = 2*Sepal.Length): Makes new variable(s) and drops existing ones.
Recommended for you
This section contains best data science and self-development resources to help you on your path.
Source: https://www.datanovia.com/en/lessons/compute-and-add-new-variables-to-a-data-frame-in-r/
0 Response to "How to Add Factors to Continuous Variable in R Dplyr by Mean"
Postar um comentário